import pandas as pd
import numpy as np
import visions as v
from visions.typesets import CompleteSet
from visions.functional import infer_type, cast_to_inferred, compare_detect_inference_frame
%matplotlib inline
The Basics
visions is a library for deterministic evaluation of semantic data types. Under normal conditions, particularly when working with unprocessed data, the type of a series is defined by it’s physical representation in memory, so although it’s trivially obvious to a practitioner that the sequence [1.0, 2.0, 3.0] should be integer, to the machine those are all floats. Here we will demonstrate how you can use visions to easily infer the semantic representations of your data for this and other more complicated data types.
# Load dataset
df = pd.read_csv(
"https://raw.githubusercontent.com/datasciencedojo/datasets/master/titanic.csv"
)
df.head()
| PassengerId | Survived | Pclass | Name | Sex | Age | SibSp | Parch | Ticket | Fare | Cabin | Embarked | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 0 | 3 | Braund, Mr. Owen Harris | male | 22.0 | 1 | 0 | A/5 21171 | 7.2500 | NaN | S |
| 1 | 2 | 1 | 1 | Cumings, Mrs. John Bradley (Florence Briggs Th... | female | 38.0 | 1 | 0 | PC 17599 | 71.2833 | C85 | C |
| 2 | 3 | 1 | 3 | Heikkinen, Miss. Laina | female | 26.0 | 0 | 0 | STON/O2. 3101282 | 7.9250 | NaN | S |
| 3 | 4 | 1 | 1 | Futrelle, Mrs. Jacques Heath (Lily May Peel) | female | 35.0 | 1 | 0 | 113803 | 53.1000 | C123 | S |
| 4 | 5 | 0 | 3 | Allen, Mr. William Henry | male | 35.0 | 0 | 0 | 373450 | 8.0500 | NaN | S |
Plotting
If you don’t already have gcc compiled you might need to install pygraphviz directly. Visions relies upon pygraphviz to visualize the relationship graph constructed between types.
Let’s make sure pygraphviz is installed so that
!conda install pygraphviz -y
Collecting package metadata (repodata.json): done
Solving environment: done
==> WARNING: A newer version of conda exists. <==
current version: 4.8.1
latest version: 4.8.4
Please update conda by running
$ conda update -n base -c defaults conda
# All requested packages already installed.
visions Building Blocks
Within visions there are two main ideas we need to get started.
Types
The first is, unsurprisingly, a Type. Think of these as literally the semantic meaning of a sequence. A String type represents the type for all sequences of exclusively strings (and the missing value indicator np.nan), a DateTime type the type for all sequences of Timestamps or Datetimes, and so on.
Typesets
The second is the Typeset, which represents a collection of types. Now that we’ve moved beyond the realm of physical types it’s possible to imagine conflicting notions of a sequences data type. For example, some users might be interested in probabilities defined as any sequence of numbers between 0 and 1 while others might instead consider them bound between 0 and 100. In order to resolve this conflict, users can encapsulate their preferred types into a single object, the Typeset. So long as each type follows a few simple rules, visions will automatically determine the relationships between types to construct a traversible relationship graph between each.
typeset = CompleteSet()
typeset.plot_graph()
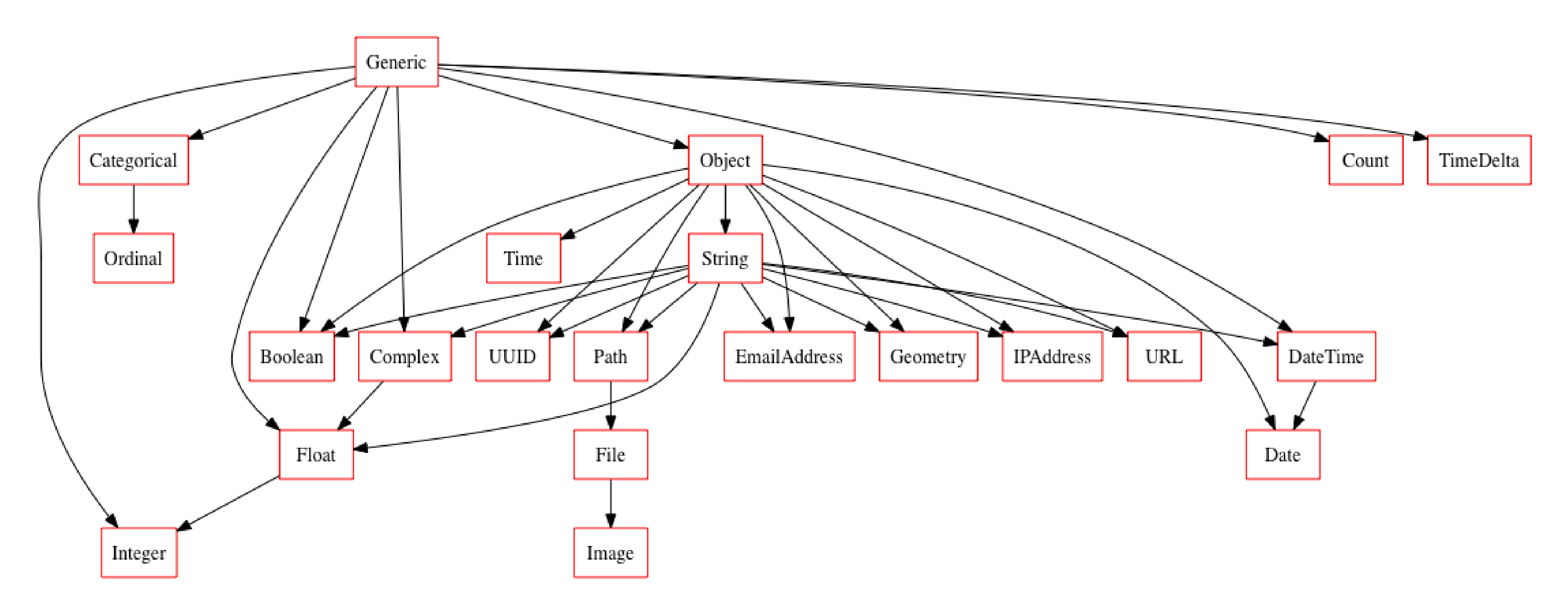
Using a Typeset
Now that we have the basic building blocks down, let’s start using our typeset. Our first task is to see visions type inference capabilities in action. In order to make the problem a little more interesting let’s also convert all of our data to strings and see what visions does.
inferred_types = compare_detect_inference_frame(df.astype(str), typeset)
pd.DataFrame({name: [data[i] for data in inferred_types]
for i, name in enumerate(['Columns', 'OriginalType', 'Inferredtype'])})
| Columns | OriginalType | Inferredtype | |
|---|---|---|---|
| 0 | PassengerId | String | Integer |
| 1 | Age | String | Float |
| 2 | Fare | String | Float |
| 3 | Survived | String | Integer |
| 4 | Embarked | String | String |
| 5 | Cabin | String | String |
| 6 | Name | String | String |
| 7 | Parch | String | Integer |
| 8 | Ticket | String | String |
| 9 | Sex | String | String |
| 10 | Pclass | String | Integer |
| 11 | SibSp | String | Integer |
Great, as we can see, visions is able to exploit the relationships instrinsic to a Typeset to infer the semantic types of our data regardless of it’s initial physical representation.
We can go a little further though and have visions losslessly coerce our data into it’s most semantically meaningful representation without our lifting a finger.
cast_df = cast_to_inferred(df, typeset)
cast_df.head()
| PassengerId | Survived | Pclass | Name | Sex | Age | SibSp | Parch | Ticket | Fare | Cabin | Embarked | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 0 | 3 | Braund, Mr. Owen Harris | male | 22.0 | 1 | 0 | A/5 21171 | 7.2500 | NaN | S |
| 1 | 2 | 1 | 1 | Cumings, Mrs. John Bradley (Florence Briggs Th... | female | 38.0 | 1 | 0 | PC 17599 | 71.2833 | C85 | C |
| 2 | 3 | 1 | 3 | Heikkinen, Miss. Laina | female | 26.0 | 0 | 0 | STON/O2. 3101282 | 7.9250 | NaN | S |
| 3 | 4 | 1 | 1 | Futrelle, Mrs. Jacques Heath (Lily May Peel) | female | 35.0 | 1 | 0 | 113803 | 53.1000 | C123 | S |
| 4 | 5 | 0 | 3 | Allen, Mr. William Henry | male | 35.0 | 0 | 0 | 373450 | 8.0500 | NaN | S |
cast_df.dtypes
PassengerId int64
Survived int64
Pclass int64
Name object
Sex object
Age float64
SibSp int64
Parch int64
Ticket object
Fare float64
Cabin object
Embarked object
dtype: object
As an aside, utilities like cast from visions functional API are simple redirections of an object oriented API defined on the typeset. Feel free to use whichever is most appealing to your personal style.
Custom Typing
This is all well and good but often we are interested in defining our own custom semantic associations with our data whether as part of an EDA pipeline, a data validation test, or something else altogether and with visions defining those types is easy.
Let’s create a BiologicalSex type corresponding to the same field in our data which has only two unique values: male and female.
from visions.types import VisionsBaseType
class BiologicalSex(VisionsBaseType):
@classmethod
def contains_op(cls, series):
return series.isin({'male', 'female'}).all()
@classmethod
def get_relations(cls):
return []
As we can see, defining a new type is easy, requiring only two methods: contains_op and get_relations. The contains op is a test to determine whether a provided series is of the semantic type represented by the class. In this case, is the data comprised exclusively of the strings male and female.
This simple definition builds a friendly API to work with data. For example, if we wanted to determine whether a sequence was of the type BiologicalSex we can do the following
df.Sex in BiologicalSex
True
df.Age in BiologicalSex
False
What about get_relations though? This method allows us to create relationships between other semantic data types. In this case, our definition of BiologicalSex assumes that all data is a String so let’s now define a relationship between the two and create a new typeset with BiologicalSex.
from visions.relations import IdentityRelation, InferenceRelation
from visions.types import String
class BiologicalSex(VisionsBaseType):
@classmethod
def contains_op(cls, series):
return series.isin({'male', 'female'}).all()
@classmethod
def get_relations(cls):
return [IdentityRelation(cls, String)]
new_typeset = typeset + BiologicalSex
new_typeset.plot_graph()
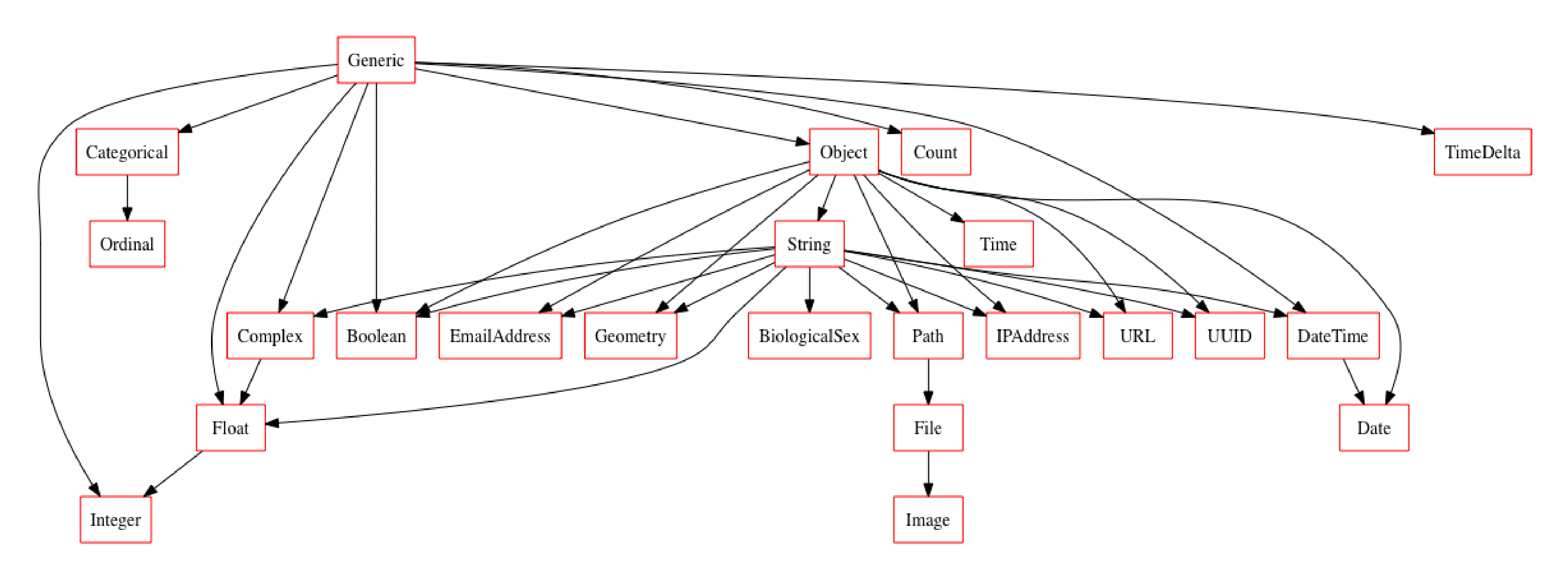
Voila, like that we have a new type appropriately inserted in the relationship graph of our new typeset and we can do all the same work done above.
Dealing with Nulls
We do have one potential problem though, we haven’t accounted for any missing value indicators in our contains_op, that means data with missing values won’t be considered to be BiologicalSex. There are numerous ways to resolve this issue (should you wish) but visions offers utility functionality to easily define nullable contains operations.
We can even use both in the same typeset by appropriately defining relations.
null_biosex_series = pd.Series([np.nan, 'male', 'female'])
null_biosex_series in BiologicalSex
False
from visions.utils.series_utils import nullable_series_contains
class NullBiologicalSex(VisionsBaseType):
@classmethod
@nullable_series_contains
def contains_op(cls, series):
return series.isin({'male', 'female'}).all()
@classmethod
def get_relations(cls):
return [IdentityRelation(cls, String)]
class BiologicalSex(VisionsBaseType):
@classmethod
def contains_op(cls, series):
return series.isin({'male', 'female'}).all()
@classmethod
def get_relations(cls):
return [IdentityRelation(cls, NullBiologicalSex, relationship=lambda s: not s.hasnans)]
null_biosex_series in NullBiologicalSex
True
Bringing it all together
new_typeset = typeset + NullBiologicalSex + BiologicalSex
new_typeset.plot_graph()
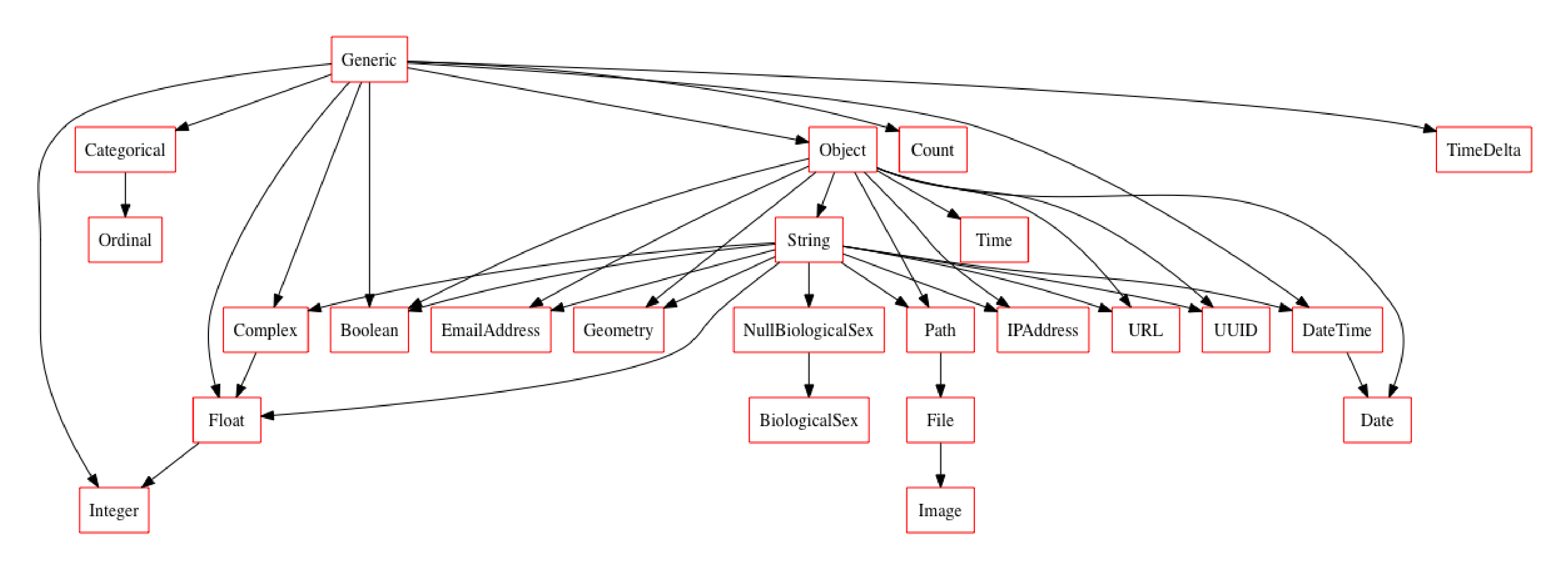
import random
new_df = df.astype(str)
new_df['NullSex'] = [random.choice(['male', 'female', np.nan]) for i in range(df.shape[0])]
inferred_types = compare_detect_inference_frame(new_df, new_typeset)
pd.DataFrame({name: [data[i] for data in inferred_types]
for i, name in enumerate(['Columns', 'OriginalType', 'Inferredtype'])})
| Columns | OriginalType | Inferredtype | |
|---|---|---|---|
| 0 | PassengerId | String | Integer |
| 1 | Age | String | Float |
| 2 | Fare | String | Float |
| 3 | NullSex | NullBiologicalSex | NullBiologicalSex |
| 4 | Survived | String | Integer |
| 5 | Embarked | String | String |
| 6 | Cabin | String | String |
| 7 | Name | String | String |
| 8 | Parch | String | Integer |
| 9 | Ticket | String | String |
| 10 | Sex | BiologicalSex | BiologicalSex |
| 11 | Pclass | String | Integer |
| 12 | SibSp | String | Integer |